Teaching

We think that genomic technologies and especially the associated computational and statistical skills are a crucial part of modern life science curricula. We try to accommodate this at all levels in our teaching. You can find all our taught courses at the course catalogue (LSF). For general information for students and curricula see the website of our faculty.
Bachelor
Wir sind (mit-)verantwortlich für folgende Kurse:
Wintersemester
- Vorlesung: Allgemeine Biologie: Prinzipien - Forschungsfelder – Geschichte (Ansprechpartner)
- Vorlesung: Grundlagen der Molekularbiologie
- Vorlesung: Molekularbiologie
- Vorlesung: "Biologie für Studierende der Zahnmedizin" (Ansprechpartner)
- Übung: Computer- und Programmierkenntnisse
- Alte Studienordnung: P 12 Modul "Humanbiologie 1": (Ansprechpartner);
nur noch als Moodlekurs verfügbar;
Praktische Übung wird durch Methoden der Tier- und Humanphysiologie ersetzt;
Klausur und Wiederholungsklausur wird nach wie vor geschrieben
Sommersemester
- Vorlesung: Tier- und Humanphysiologie
- Vorlesung und Übung: "Pretty Plots - Visualisierung statistischer Daten" (berufsqualifizierend)
- Vorlesung: Grundlagen der Biologie, Teil 2 (Für Geowissenschaftler und Bioinformatiker, wird von der TUM organisiert)
Master
We are contributing to the following master programs:
We are (co-)responsible for the following courses:
NOTE: PRACTICALS ARE CENTRALLY DISTRIBUTED!
So just enrol in the LSF and attend the central distribution meeting!
Winter semester
- Lecture "Human genomics I"
- Seminar "Current topics in Statistical Genomics" (especially for advanced students and/or students doing their theses/research course in our lab)
- Seminar and Practical course "Computational analysis of RNA-Seq data"
- Practical course "Computational biology in molecular and cellular biology" (R-week).
![]()
These classes are supported by DataCamp, the most intuitive learning platform for data science. Learn R, Python and SQL the way you learn best through a combination of short expert videos and hands-on-the-keyboard exercises. Take over 100+ courses by expert instructors on topics such as importing data, data visualization or machine learning and learn faster through immediate and personalised feedback on every exercise.
Summer semester
- Lecture "Genomics of Human Diseases"
- Seminar "Current topics in Statistical Genomics" (especially for advanced students and/or students doing their theses/research course in our lab)
- Practical course "Pretty plots"
Further information is available on the faculty website on Master Programs as well as in the course catalogue (LSF).
Master Molecular and Cellular Biology
Please refer to the LSF and the curriculum overview for possible modules. For students interested in our teaching activity we suggest the following modules (many other options exist though):
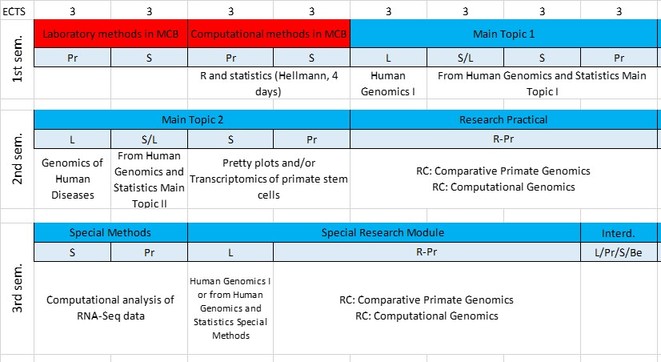
red: required courses, blue: elective courses.
Pr: practical course, S: seminar, L: lecture, R-Pr: research project, D: disputation, C: colloquium, Be: vocational course.
Research Courses
Please consult the guidelines for research courses available here.
Internal Research Courses
In general, we expect that RC or IRT (Master EES) applicants have visited our lectures, seminars and/or our practical master courses. We offer two types of RCs:
- Comparative Primate Genomics:
please apply to enard@bio.lmu.de; this involves RCs that are mainly wet lab and involve our research of Foxp2, cancer evolution, primate iPSCs and (single-cell) RNA-seq - Computational Genomics:
please apply to hellmann@bio.lmu.de; this involves RCs that are computational and involve our research of cancer evolution, regulatory evolution and population genomics; programming skills and/or statistical skills are mandatory and hence participation in Pretty Plots and/or Computational analysis of RNA-Seq data.
When applying please provide your CV, your record of transcripts, a short description of your motivation and the approx. timeframe you would be able to do your RC.
External Research Courses
Please consult our guidelines for external research courses available here.
Bachelor and Master Theses
Our bachelor theses projects are presented at the annual bachelor fair in December.
For master theses we generally expect people to have participated in our courses and have done a Research course in our lab. We offer computationally oriented projects (contact Dr. Hellmann, hellmann@bio.lmu.de) and experimentally oriented projects (contact Professor Dr. Enard, enard@bio.lmu.de).

